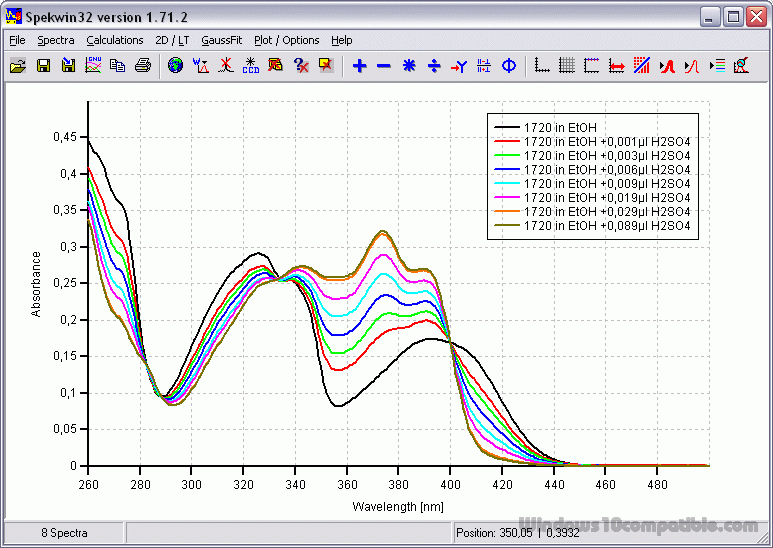
Algorithms that estimate secondary structure depend on correct protein concentrations.Over-estimating or under-estimating the protein concentration interferes with signal.An accurate protein concentration is required for all CD experiments.DMSO, formamides, and imidazole absorb strongly in the far UV region, and thus should be avoided in a CD experiment.Triton detergents should be avoided, as they can oxidize readily and form UV-absorbing materials. Some detergents are fairly transparent in the far UV (e.g.DTT, ß-ME, or EDTA can be present at low concentrations (1 mM).Chloride ions will result in a loss of signal below about 200±5 nm, which affects secondary structure estimation, but generally will not block detection of the alpha-helical peaks (208 nm and 222 nm) or the beta-sheet peak (218 nm).Potassium fluoride may be a good substitute for chloride-containing salts. 
If salt is required, SO 4 2- or F - are preferred counter ions, as they are transparent in the far UV.NaCl and Tris buffers are not recommended but can be tolerated at low concentrations.Ideal aqueous solutions contain as little chloride as possible.Phosphate buffers with little-to-no NaCl are recommended.Ensure that the buffer blank is well matched to the final dilution of your protein (even trace amounts of some solvents will interfere with the CD measurements).Always take a scan of your buffer blank to determine whether absorbance interferes with your region of analysis.with no buffer absorbance through the range of the CD spectrum.in which your protein is well behaved and soluble.The ideal CD experiment is performed in a buffer:.Many commonly used buffers and additives absorb in the far UV region used for CD measurements. Solvent absorbance can severely interfere with the CD signal. ValiDichro tests the quality of CD data based on based on common characteristics of CD protein spectra observed in the literature.īuffer selection is critical for accurate CD measurements. The YouTube channel provides video tutorials on a variety of CD-related topics, including the successful measurement of a CD spectrum and analysis using DichroWeb and CDToolX.īeStSel (Beta Structure Selection), a free online tool based on a novel method for the secondary structure determination and fold recognition from protein circular dichroism spectra, especially beta-sheet rich proteins. Protein Circular Dichroism Data Bank, a database of CD data Jasco Training Videos provide more information on various topics related to CD measurements and the Spectra Manager program.ĭichroWeb , a free online tool for determining protein secondary structures based on CD spectra.Īcademic users may apply for a username and password.ĬDToolX, a free, downloadable software program that enables processing, displaying, archiving, calibrating, comparisons, and analyses of circular dichroism (CD) spectroscopic data. J-815 RETIRED APRIL 2023 - CMI Jasco J-815 CD Getting Started Guide. Instrument and software manuals are located in the “Jasco Manuals” desktop folder on the instrument computer. It does not store any personal data.CMI Jasco J-1500 CD Getting Started Guide, guide to experimental design and standard Far-UV CD protocols.Īdditional resources are available at the instrument. The cookie is set by the GDPR Cookie Consent plugin and is used to store whether or not user has consented to the use of cookies. It allows the website owner to implement or change the website's content in real-time. 
This cookie is used by the website's WordPress theme. 
It works only in coordination with the primary cookie. Records the default button state of the corresponding category & the status of CCPA. The cookies is used to store the user consent for the cookies in the category "Necessary". This cookie is set by GDPR Cookie Consent plugin. The cookie is used to store the user consent for the cookies in the category "Analytics". These cookies ensure basic functionalities and security features of the website, anonymously. Necessary cookies are absolutely essential for the website to function properly.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |



 RSS Feed
RSS Feed
